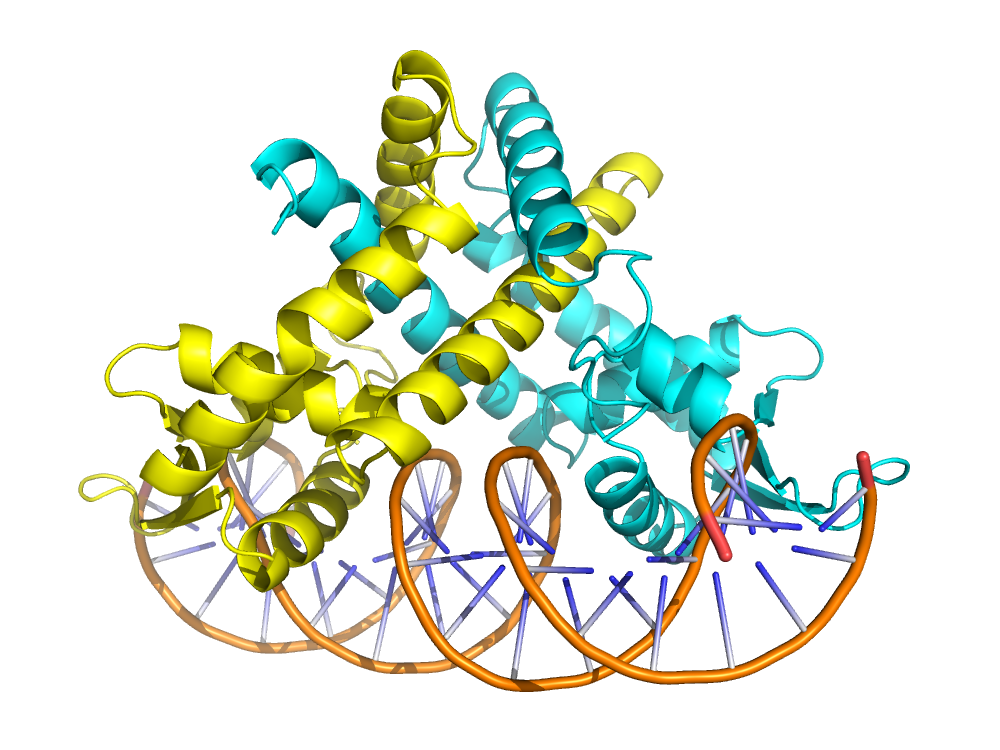
This example shows how to filter PDB by protein DNA complexes
In [1]:
from pyspark import SparkConf, SparkContext
from mmtfPyspark.io import mmtfReader
from mmtfPyspark.filters import ContainsDnaChain, ContainsLProteinChain
from mmtfPyspark.structureViewer import view_structure
In [2]:
conf = SparkConf().setMaster("local[*]") \
.setAppName("FilterProteinDnaComplexDate")
sc = SparkContext(conf = conf)
In [3]:
path = "../../resources/mmtf_reduced_sample/"
pdb = mmtfReader.read_sequence_file(path, sc)
In [4]:
structures = pdb.filter(ContainsLProteinChain()) \
.filter(ContainsDnaChain())
In [5]:
count = structures.count()
print(f"Number of entires that contain L-protein and L-DNA: {count}")
Number of entires that contain L-protein and L-DNA: 124
In [7]:
structure_names = structures.keys().collect()
view_structure(structure_names)
Out[7]:
<function mmtfPyspark.structureViewer.view_structure.<locals>.view3d>
In [6]:
sc.stop()