This demo shows how to create a dataset of ATP Interating atoms.
In [1]:
from pyspark import SparkConf, SparkContext, SQLContext
from mmtfPyspark.datasets import groupInteractionExtractor
from mmtfPyspark.io import mmtfReader
from mmtfPyspark.webfilters import Pisces
import time
In [2]:
conf = SparkConf().setMaster("local[*]") \
.setAppName("ATPInteractionAnalysisDemo")
sc = SparkContext(conf = conf)
In [3]:
path = "../../resources/mmtf_full_sample/"
pdb = mmtfReader.read_sequence_file(path, sc)
In [4]:
seqId = 40
resolution = 2.0
pdb = pdb.filter(Pisces(seqId, resolution))
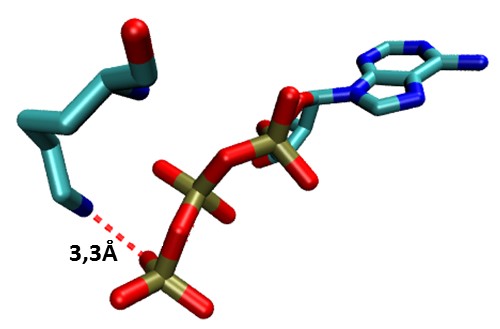
ATPInteraction
In [5]:
finder = groupInteractionExtractor("ATP", 3)
interactions = finder.get_dataset(pdb).cache()
In [6]:
interactions = interactions.filter("atom1 LIKE('O%G')")
In [7]:
interactions.printSchema()
interactions.show(20)
root
|-- structureId: string (nullable = false)
|-- residue1: string (nullable = false)
|-- atom1: string (nullable = false)
|-- element1: string (nullable = false)
|-- index1: integer (nullable = false)
|-- residue2: string (nullable = false)
|-- atom2: string (nullable = false)
|-- element2: string (nullable = false)
|-- index2: integer (nullable = false)
|-- distance: float (nullable = false)
+-----------+--------+-----+--------+------+--------+-----+--------+------+---------+
|structureId|residue1|atom1|element1|index1|residue2|atom2|element2|index2| distance|
+-----------+--------+-----+--------+------+--------+-----+--------+------+---------+
| 1B0U| ATP| O3G| O| 261| SER| OG| O| 36|2.6808183|
| 1B0U| ATP| O3G| O| 261| HOH| O| O| 272|2.7428646|
| 1B0U| ATP| O1G| O| 261| HOH| O| O| 293| 2.750308|
| 1B0U| ATP| O2G| O| 261| HOH| O| O| 299|2.8596847|
| 1B0U| ATP| O3G| O| 261| HOH| O| O| 356|2.6914153|
+-----------+--------+-----+--------+------+--------+-----+--------+------+---------+
In [8]:
n = interactions.count()
print(f"Number of interactions: {n}")
Number of interactions: 5
In [9]:
topGroups = interactions.groupBy("residue2").count()
topGroups.sort("count", ascending = False).show(10) # Sort descending by count
+--------+-----+
|residue2|count|
+--------+-----+
| HOH| 4|
| SER| 1|
+--------+-----+
In [10]:
topGroupsAndAtoms = interactions.groupBy("residue2","atom2").count()
topGroupsAndAtoms.withColumn("frequency", topGroupsAndAtoms["count"] / n)\
.sort("frequency", ascending = False) \
.show(10)
+--------+-----+-----+---------+
|residue2|atom2|count|frequency|
+--------+-----+-----+---------+
| HOH| O| 4| 0.8|
| SER| OG| 1| 0.2|
+--------+-----+-----+---------+
In [11]:
sc.stop()